Single-cell multi-omics characterize colorectal tumors, adjacent healthy tissue and matched (tumor) organoids identifying CRC-unique features
Abstract
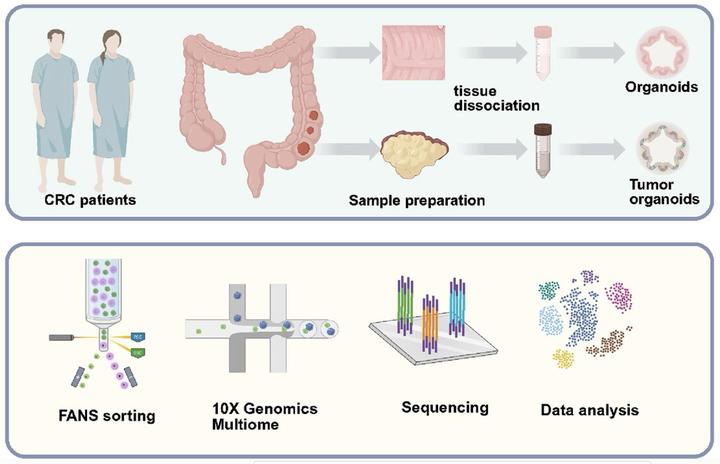
Colorectal cancer (CRC) arises in the colorectal tissue driven by genetic disorder or the accumulation of somatic mutations, leading to abnormal epithelial cell growth. In this study, we employed single-nucleus multi-omics analysis, including single-nucleus RNA-seq and single-nucleus ATAC-seq, on over 100,000 high-quality nuclei to investigate the molecular landscape of both primary tissue and patient-derived organoids (PDOs). Our analysis showed that normal PDOs (N-PDOs) derived from tissue adjacent to tumors replicate the cellular composition and differentiation trajectory of colorectal crypts. In contrast, tumor PDOs (T-PDOs) showed patient-specific transcriptomic and epigenomic heterogeneity yet consistently maintained a stem cell-like state. T-PDOs retained the somatic mutation profile of the primary tumor while also exhibiting de novo mutations not detected in either the primary tumor or N-PDOs. Notably, inferred cell–cell interaction analysis highlighted the activin signaling pathway as a potential unique feature of fibroblast-epithelial interactions within the tumor microenvironment. This study provides a comprehensive view of the transition from normal to malignant colorectal epithelium and underscores the utility of PDOs as a faithful model for capturing both conserved and patient-specific features of colorectal cancer.